Research
Genomic Insights Into the Origin and Diversification of Diatoms
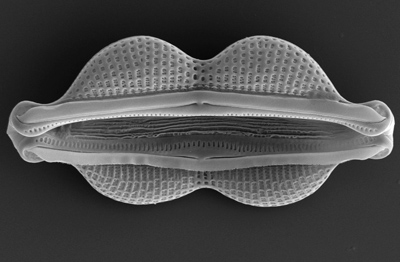 With some 200,000 species, diatoms are probably the most species-rich lineage of microbial eukaryotes. In sharp contrast, the sister group to diatoms (the Bolidophyceae) includes fewer than 20 species. Comparing bolidophyte genomes to those of early diverging diatoms will allow us to pinpoint the key genomic changes that took place in the common ancestor of all diatoms. We're sequencing the genomes of diatom species that bracket the earliest splits in the phylogeny and comparing them to the genomes of closely related outgroup species. These data will provide key insights into the origins of novel diatom traits, including those that may have facilitated their nearly unprecedented phylogenetic and ecological diversification. Image credit: Elizabeth Ruck.
With some 200,000 species, diatoms are probably the most species-rich lineage of microbial eukaryotes. In sharp contrast, the sister group to diatoms (the Bolidophyceae) includes fewer than 20 species. Comparing bolidophyte genomes to those of early diverging diatoms will allow us to pinpoint the key genomic changes that took place in the common ancestor of all diatoms. We're sequencing the genomes of diatom species that bracket the earliest splits in the phylogeny and comparing them to the genomes of closely related outgroup species. These data will provide key insights into the origins of novel diatom traits, including those that may have facilitated their nearly unprecedented phylogenetic and ecological diversification. Image credit: Elizabeth Ruck.
Collaborator: Norm Wickett (Chicago Botanic Garden)
The Genomic Basis of Microbial Adaptation to New and Changing Environments
Although evolutionary biology is a historical science, it provides a powerful means of looking forward as well—offering predictions on issues ranging from the spread of human disease to adaptation of species to global climate change. Increased atmospheric carbon and global warming are having profound impacts on the world’s oceans. Changing precipitation patterns and melting polar ice caps are freshening large regions of the ocean, resulting in changes in water stratification, convection, and nutrient availability. Although these changes are predicted to have important impacts on the phytoplankton communities in these areas, relatively few data are available to predict how phytoplankton—which produce 40% of the earth’s oxygen and form the base of marine food webs—will respond to a rapidly changing ocean. Laboratory experiments have shown that microbial eukaryotes can adapt quickly to environmental change in vitro, but it’s not always clear how these results extrapolate to natural systems. This project leverages a natural evolutionary experiment, the colonization of low-salinity habitats in the Baltic Sea by an ancestrally marine diatom, Skeletonema marinoi, to determine the mechanisms and rate of adaptation to environmental stress across ecological and evolutionary timescales. Comparative and population genomic data, together with experimental transcriptomic data, will be combined to determine the tempo and mode of environmental adaptation of S. marinoi populations spanning the North–Baltic Sea salinity cline. Data from a related species, Skeletonema potamos, which has independently adapted to freshwaters, will show whether there is more than one adaptive solution to managing salinity stress, providing general insights into the adaptive potential of phytoplankton to a rapidly changing ocean.
Collaborators: Teo Nakov (University of Arkansas), Matt Hahn and Geoffrey House (Indiana University), Anna Godhe and Mats Töpel (University of Gothenburg), Anke Kremp (Finnish Environment Institute)
The Evolutionary Transition Between Marine and Fresh Waters
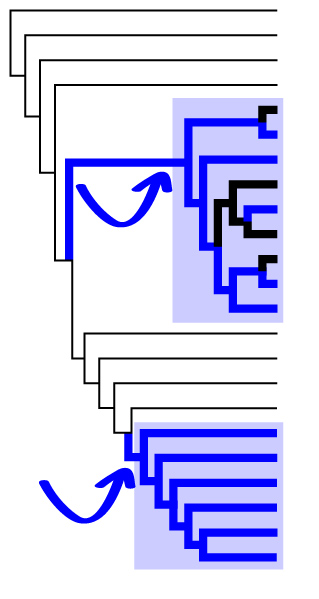 Freshwater colonizations are landmark events in diatom evolution, having led to the origins of perhaps tens of thousands of species. A primary goal of this research is to understand the morphological, physiological, and genetic adaptations that have allowed diatoms to diversify into freshwater environments, which present numerous obstacles to marine colonists. We're using a blend of phylogenetic, experimental, and genomic approaches to understand how diatoms have successfully, and repeatedly, conquered "the salinity barrier". This project is focused primarily on Thalassiosirales, a lineage that presents an ideal model system for several reasons: (1) most species are easily cultured; (2) the lineage includes the model species, Thalassiosira pseudonana, which offers excellent genomic resources, and; (3) there is a strongly supported phylogenetic hypothesis, laying out a rich and well-resolved historical pattern of marine–freshwater transitions. This history includes numerous successful freshwater colonizations, so nature has played out this experiment multiple times. Our goal is to determine whether these natural experiments played out in the same, or different, ways.
Freshwater colonizations are landmark events in diatom evolution, having led to the origins of perhaps tens of thousands of species. A primary goal of this research is to understand the morphological, physiological, and genetic adaptations that have allowed diatoms to diversify into freshwater environments, which present numerous obstacles to marine colonists. We're using a blend of phylogenetic, experimental, and genomic approaches to understand how diatoms have successfully, and repeatedly, conquered "the salinity barrier". This project is focused primarily on Thalassiosirales, a lineage that presents an ideal model system for several reasons: (1) most species are easily cultured; (2) the lineage includes the model species, Thalassiosira pseudonana, which offers excellent genomic resources, and; (3) there is a strongly supported phylogenetic hypothesis, laying out a rich and well-resolved historical pattern of marine–freshwater transitions. This history includes numerous successful freshwater colonizations, so nature has played out this experiment multiple times. Our goal is to determine whether these natural experiments played out in the same, or different, ways.


